Zoo/PhytoImage analyzes plankton samples through image analysis and machine learning. Plankton images from various origins can be used.
Analyze your plankton through digitized images
Plankton samples are traditionally collected with plankton nets and analyzed by taxonomists using a stereomicroscope and determination keys. This work is labor-intensive and requires highly-specialized and well-trained people.
With the advances in image analysis and machine learning, the analysis of digitized plankton images can support, ease and speed-up the work. The free Zoo/PhytoImage software (licensed under GNU/GPL) provides a complete solution for such a job.
Download
Only odd version numbers are public. Even version number are for internal development only.
-
Zoo/PhytoImage v.5 (latest)
Version 5 introduces the ‘tinted vignettes’ that ease to spot the boundaries of the particles. Analysis is still done in grayscale, but the colored vignette is obtained by mixing the mask, the OD-calibated image and the visually-enhanced one into one picture. Each item can also be re-extracted, and the mask + OD images can be used to reanalyze the item.
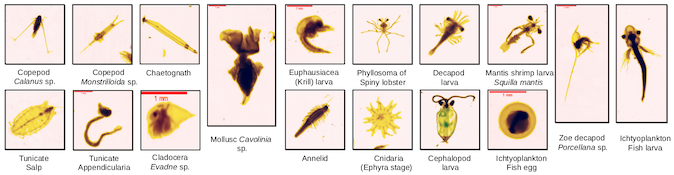
-
Zoo/PhytoImage v.3
Version 3 proposes a new storage format for zooimage samples (.zidb) that is much more efficient than the previous .zid format.
-
Zoo/PhytoImage v.1
The original version!
-
Github repository
-
Manual
-
FlowCAM example
-
Macrophotography example
-
Idem (training set)
-
Microphotography ewample
-
Idem (training set)
-
Scanner example (color 24bit)
-
Idem (training set) 13 Scanner example (gray 16bit)
-
Idem (training set)
Install
TODO…
Get help
After downloading the sofware, look at the Zoo/PhytoImage user’s manual that explains in details how to install the software and how to use it (quick tutorial). Additional example datasets should be used for training before working with your actual data. If you have further, or specific questions, you should use the dedicated forum.

